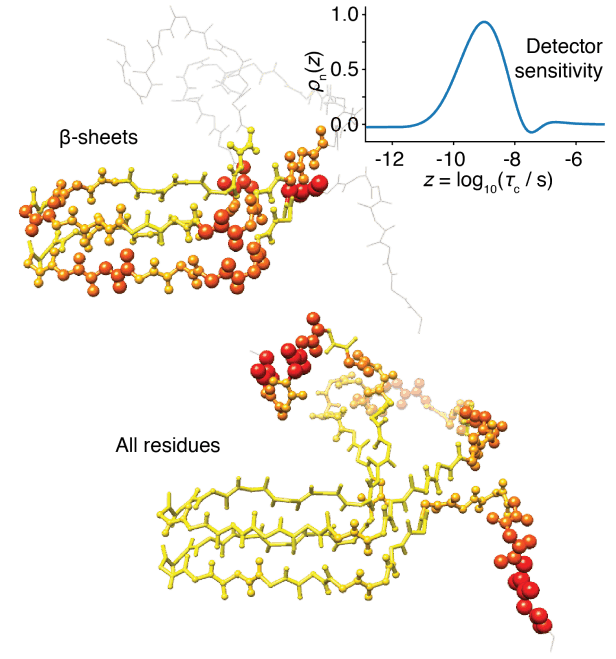
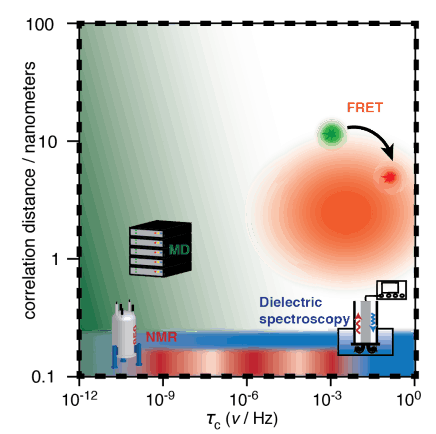
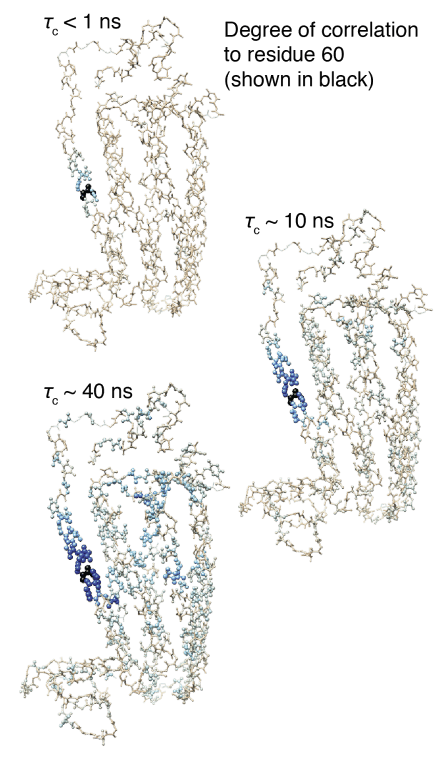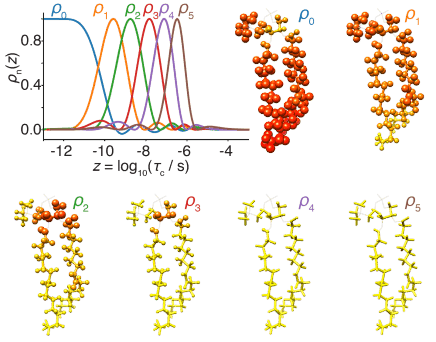Detectors The "detector" method of analyzing dynamics has been recently developed, as a means of reducing bias introduced by the analysis of NMR relaxation data using explicit models. Detectors are being further developed also as a means of analyzing molecular dynamics simulation. This includes comparison of dynamic behavior between experiment and simulation and correlation analysis between different sites of a protein, in order to connect local and overall motion.
Figure: MD-derived detector responses for
HET-s(218-289) fibrils near 1 ns